Compositional Systems Biology
Compositional Systems Biology is our approach to building infrastructure for integrative biological modeling: a generalizable architecture for composing multiscale simulations from modular models with transparent data flow.
Our infrastructure development focuses on:
- Composition Framework: a standard for connecting heterogeneous processes through modular interfaces and orchestration patterns.
- Composition Schema: a unified format for defining models, data structures, and process interactions, supporting reproducibility and automated composition.
- Compositional Software: open-source tools for simulation runtime, schema validation, model annotation, and API-based deployment.
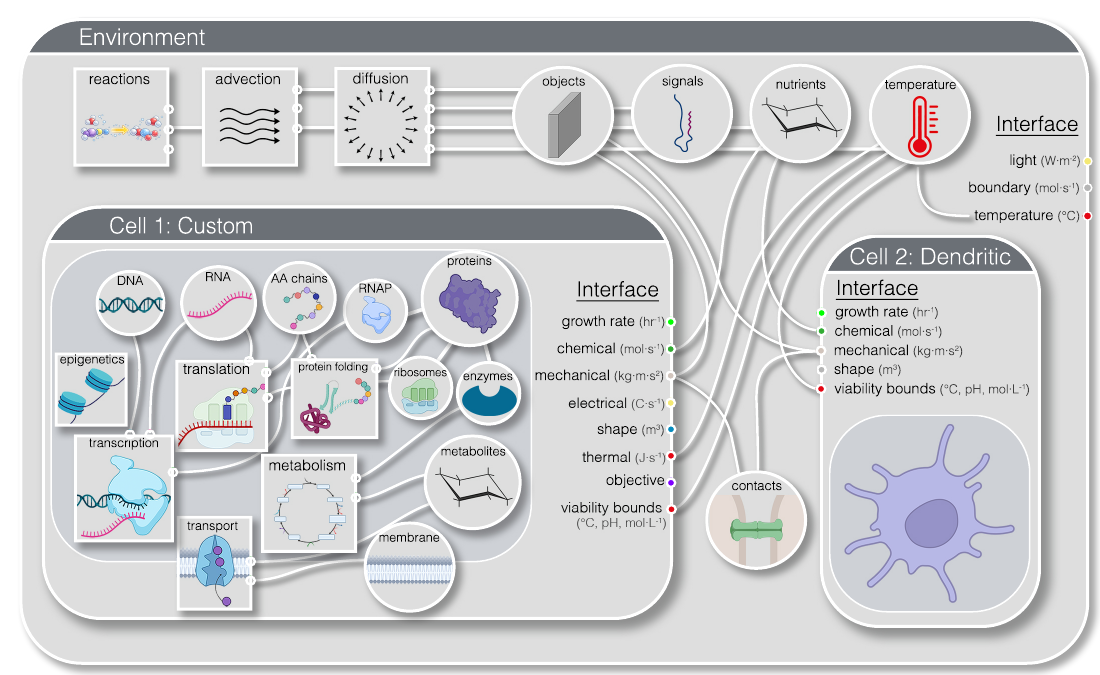
- Biological Interface Definitions: standard schemas for cells, molecules, and environments, keeping simulations consistent and extensible.
- Collaborative Modeling Infrastructure: shared registries that let research groups reuse modular components across domains.
The goal is to reduce the friction of building and sharing complex biological models, creating an open ecosystem where multiscale simulations evolve through contributions from many research groups.
This infrastructure is being built as Vivarium 2.0, a redesign of the Vivarium framework around the process bigraph formalism. It introduces a typed schema system and composition engine that make models easier to define, validate, compose, and visualize. The core software is developed openly across process-bigraph (the composition engine), bigraph-schema (the type and schema system), and bigraph-viz (visualization of composite models).
The approach rests on a formal foundation: process bigraphs give a rigorous mathematical account of how heterogeneous processes share state and compose, keeping combined models well-defined. See the Process Bigraph Supplemental Materials for the formal treatment.
E. coli Whole-cell Model
We are developing a mechanistic model of E. coli that integrates gene regulation, metabolism, and cellular physiology across molecular, cellular, and population scales. With over 19,000 parameters, it predicts gene expression, metabolite levels, protein interactions, and whole-cell properties such as biomass and growth rate.
We use the model as a discovery tool to:
- Complete functional annotations by predicting the roles of uncharacterized genes from simulation–data comparisons.
- Capture population heterogeneity by simulating variation in growth and physiology across thousands of cells.
- Model environmental context by testing how nutrients and stressors shape adaptation and behavior.
- Enable simulation-based inference using probabilistic and ML methods for parameter estimation and experimental design.
By unifying diverse data in an extensible simulation, the model supports systems-level reasoning and discovery in microbial physiology.
Ocean Microbiome
We connect genome-scale metabolic models of ocean bacteria and plankton to biogeochemical models of the water column, quantifying how microbial activity affects climate. This work includes:
- Metabolic Modeling: detailed models of microbial metabolism in ocean bacteria and plankton.
- Biogeochemical Integration: linking these models to water-column cycles to assess nutrient and carbon fluxes.
- Climate Impact: quantifying how microbial carbon sequestration and nutrient cycling influence climate.
This work is conducted in collaboration with the Center for Chemical Currencies of a Microbial Planet at the Woods Hole Oceanographic Institution.
Gut Microbiome
Our gut microbiome research, led by PhD students and undergraduates in the lab, builds a spatial dynamical model of the human intestines. Areas include:
- Ecological Niches: mapping intestinal niches, their microbial communities, and the spatially structured environment shaped by peristaltic flow.
- Microbial Community Interactions: how microbes interact and coexist, from molecular networks to whole-gut physiology, including microbe–host interactions such as mucus production by goblet cells.
- Layered Complexity: addressing the hierarchical structure and diverse interactions of the gut microbiome.
- Dynamic Modeling: starting from stoichiometric models, then adding multi-species interactions and processes like digestion, absorption, and peristalsis.
- Multi-Omics Data Integration: incorporating multi-omics data into spatial models of the gut environment.
The goal is to integrate these datasets and uncover the microbial interactions that drive gut health and disease.
